 |
| King Cobra (Ophiophagus hannah; top) and Burmese Python (Python bivittatus; bottom), the two snake species whose genomes were fully sequenced in 2013 |
One year ago today, the first snake genomes ever sequenced hit the newsstands. OK, so two papers in Proceedings of the National Academy of Sciences isn't exactly the cover of Time magazine to most people, but it was big enough news that it was covered by The Huffington Post and the two most prominent interdisciplinary scientific journals, Science and Nature, the former devoting a special section to the event. One year later, dear reader, welcome to the Life is Short, but Snakes are Long coverage of the snake genome project. So just what is the big deal about these snake genomes anyway, and what's changed in snake biology in the year that they've been available?
In one way, sequencing a snake genome means that snakes finally join the illustrious ranks of lab animals like the mouse, rat, guinea pig, fruit fly, and amoeba, all of whom have already had their genomes sequenced. By now the genomes of several hundred species have been sequenced, starting with a virus in the 1970s, and the first archaeon, bacterium, and eukaryote within one year of one another in 1995-96. The first animal genome sequenced was that of the model nematode Caenorhabditis elegans in 1998, and the first vertebrate was a pufferfish, so chosen because its genome is so small, in 2002 (although an incomplete first draft of the human genome preceded that by a year). As of 2014, we're now up to just over 100 vertebrate species, about 60 of which have been annotated and formally published, as well as numerous other animals, plants, fungi, protists, and prokaryotes. Last week, Science highlighted drafts of 38 new bird and 3 new crocodilian genomes, the largest single release of vertebrate genomes to date. But we are still a long way from sequencing the genomes of all known species. Why have we chosen the species we have? What does it mean to sequence a genome, exactly, and why do we do it?
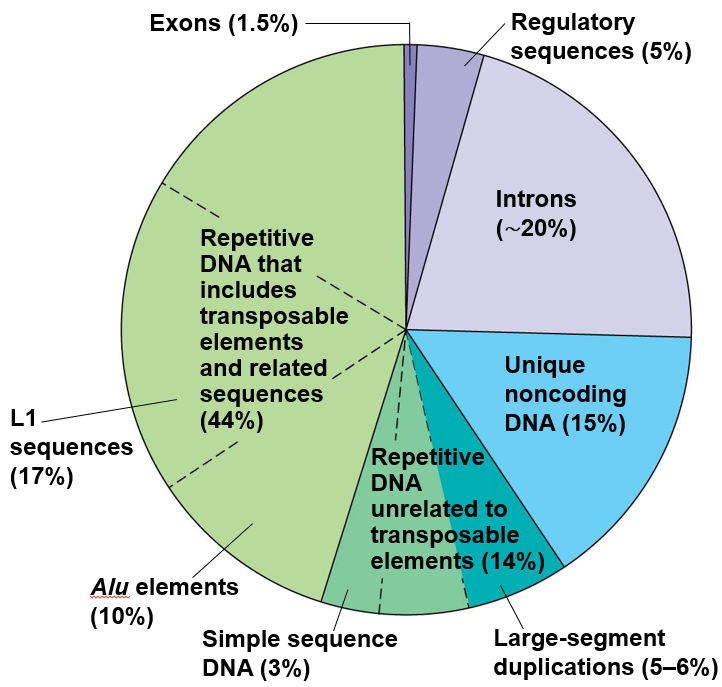 |
| Breakdown of what the human genome consists of. Exons are coding DNA. From Reece et al. (2013) |
 |
| Avian tree of life based on whole-genome sequences. We're still several years away from a tree like this for squamate reptiles. From Jarvis et al. 2014 |
3. This is different from knowing all the possible variants (often called alleles) of those genes. It's often said that a person has "the genes for" something, when in reality all people have the same genes, with different alleles. For example, if the person whose genome was sequenced in the Human Genome Project had brown eyes, we'll just have the gene sequences for brown eyes, not for blue or green. In order to get an idea of all the possible variants of all the genes in a species, we'll need to sequence the genomes of many individuals. Some genes, such as those involved in the immune system, have over 1,500 alleles (the "gene pool"), no more than two of which occur within the genome of a single individual (one from the mother and one from the father). So understanding the entire gene pool of a species is a very daunting task, given that we only have whole genomes for a few hundred species (one individual each), with multiple individuals of a few species, including humans.4 Population genomics is an emerging field, yet to be applied to snakes in any form, although apparently a few projects are in the works.
So what have we learned from these snake genomes? Here are the basics:
It's not yet clear to what extent the evolution of these novel toxic venom proteins corresponded with a shift to higher levels of their expression in the venom gland and lower levels of expression elsewhere. Although it seems obvious that their expression in non-venom-gland tissues would be harmful, their non-toxic orthologs are expressed in tissues as diverse as the kidney and brain in pythons, and no one has yet measured their expression outside of the venom gland in venomous snakes. Alternatively, gene duplication might have taken place after the change in function, if the genes in question were alternatively spliced to produce both toxic and non-toxic proteins from the same gene. Evolution of siRNA and other regulatory elements (which is hard to detect because there's still a lot we don't understand about how it works) could then restrict expression of a particular splice variant to the venom gland, which could explain why we're seeing evidence that the venom protein genes themselves are often still expressed in other tissues even though they are capable of coding for highly toxic proteins that must be maintained in the venom gland in a competent but inactive state.
The cobra genome by itself does not answer these questions, even with help from that of the python. In order to fully understand the evolution of snake venoms (with major implications for public health, particularly in developing countries, not to mention the potential of venoms to be used as drugs), we'll need genomic, transcriptomic, and proteomic data from numerous snake species.
So what have we learned from these snake genomes? Here are the basics:
- Snake genomes are about half the size of the human genome (although an organism's complexity is not directly proportional to its genome size; for example, some salamander genomes are more than 60 times larger than the human genome).
- The proportion of repetitive elements (the most common form of "junk DNA") in snake genomes is about the same as that in humans (~60%).
- Snakes have a faster baseline rate of evolution than other reptiles, birds, or mammals, as
evidenced by their larger accumulation of neutral substitutions. And colubroid snakes have rates even faster than that of snakes at large.
Red represents fast rates of neutral substitution
From supplement to Castoe et al. 2013 - Adaptive evolution (as evidenced by functional, non-neutral, changes to genes) in snakes has happened to over 500 genes, especially those involved in the development of the limbs, spine, skull, and eye, and those regulating the function of the cardiovascular system, lipid and protein metabolism, and cell birth and death. We already knew that all of these systems in snakes were highly modified relative to other vertebrates, and now we know that the genes that underlie them are too.
- Some groups of genes have grown or shrank in snakes - for example, snakes have a lot more genes coding for vomeronasal receptors, and a lot fewer genes coding for opsins, which are light-sensitive proteins in the eye. This makes sense given what we know about snake sensory systems.
- Changes to gene expression that happen after a snake feeds involve thousands of genes that control rapid changes in organ size—but genes that control cell division change in the kidney, liver, and spleen, organs that grow by cell division, but not in the heart, which grows when individual existing cells get larger.
- Snake genomes contain endogenous viral elements from three families of viruses that have recurrently infiltrated their DNA over the past 50 million years. This is actually not rare, although it is bizarre and awesome that the 'fossils' of these ancient viral genomes can be identified in their host genomes even after tens of millions of years, and it can help us better understand both the biology of viruses and that of their snake hosts, including how viruses have contributed functions to the genetic repertoires of their hosts.
- A snake has a gene that makes a protein somewhere in its body, including possibly in its salivary or venom gland5
- The gene for that protein is duplicated by accident during routine DNA replication or repair, resulting in a new, spare copy of the gene
- The effects of selection are relaxed on the duplicate gene, which gives it opportunities to mutate (because, if it does, no harm is done; the original copy continues to perform its original function)
- Mutations to transcription-factor binding sites change the signal for where the duplicate gene should be expressed, causing the new protein to be made only in the venom gland
- If the new protein helps the snake catch more prey, it improves fitness and causes natural selection
- Because the old protein is still being made, the new gene and protein are free to evolve to become more toxic or to take on some new function
- The new copy of the gene may become duplicated again, and subsequent new copies may mutate further, leading to diversification within a gene/toxin family6
 |
| The King Cobra venom gland, with expression profiles of the venom (left) and accessory gland (right). From Vonk et al. 2013 |
The cobra genome by itself does not answer these questions, even with help from that of the python. In order to fully understand the evolution of snake venoms (with major implications for public health, particularly in developing countries, not to mention the potential of venoms to be used as drugs), we'll need genomic, transcriptomic, and proteomic data from numerous snake species.
Characterization of genomic biodiversity has the potential to change our understanding of evolution in fundamental ways. From explaining how snakes are capable of physiological feats to helping us understand how new genes appear, what "junk DNA" does, and what the tree of life looks like, genome sequencing is one of the most exciting current frontiers in biology. As in many things, snakes are (one of) the last groups of vertebrates to the party (although it's worth noting that there aren't any fully annotated salamander or caecilian genomes yet). A snake genome doesn't add a whole lot to the picture of the vertebrate tree of life, because the Green Anole genome, sequenced in 2011, represents squamates on the tree, and no one is arguing that snakes aren't squamates. But, within squamates there are a number of puzzling unresolved relationships, including such fundamental questions as the origin of snakes and the placement of iguanians. In the interest of helping to shed light on these, and on the aforementioned complexity of snake venom evolution, another 10 or so snake genomes are likely to come out within the next couple of years, including those of the:
- Texas Blindsnake (Rena dulcis)
- Reticulate Wormsnake (Amerotyphlops reticulatus)
- Red Pipesnake (Anilius scytale)
- Mexican Burrowing Python (Loxocemus bicolor)
- Round Island Splitjaw Snake (or "boa"; Casarea dussumieri)
- Boa Constrictor (Boa constrictor)
- Western Diamond-backed Rattlesnake (Crotalus atrox)
- Speckled Rattlesnake (Crotalus mitchelli)
- Copperhead (Agkistrodon contortrix)
- Eastern Coralsnake (Micrurus fulvius)
- Cloudy Snail-eating Snake (Sibon nebulatus)
- Common Gartersnake (Thamnophis sirtalis)
As you can probably see if you know your snake taxonomy, these species represent a scattering of well-known snakes from each of the major branches of the snake tree. They have been strategically chosen to enable snake biologists to use them to put together a well-supported skeleton of the snake tree of life. However, several branches (such as the dwarf pipesnakes, acrochordids, and lamprophiids) are still missing.7 In particular, an atractaspidid genome would be useful in building a better understanding of the role of convergence in snake venom evolution - resolving the debate between proponents of a single ancient origin for venom and those of several more recent, independent origins. Genomes of scolecophidian blindsnakes and toxicoferan lizards such as Gila monsters will also help resolve this question. Hopefully, these genomes and others will continue to illuminate evolutionary biology for us in ways Darwin could have scarcely imagined.
1 Because genome sequences contain so much data, they are stored electronically and require a large amount of computing power and storage capacity. The computing power is actually more limiting than the biochemistry right now. A human genome contains about 6 billion base pairs (one for each person on Earth in 1999), which take up a couple of gigabytes. If that doesn't sound that impressive, imagine all that information stored in every one of your cells, then compare the size of a cell with that of a microchip here.↩
2 This is not to say that (as has been presumed by many) molecular data are inherently superior to morphological data, especially in the case of extinct fossil taxa, from which we cannot garner much molecular information (although that generalization too has been challenged).↩
3 How are the individuals whose genomes are sequenced chosen? The unsatisfying answer is that the scientists involved typically use whatever individuals are convenient. Specifically, the cobra and python genomes seem to have been taken from animals from the pet trade. We may not know the true geographic origin of these individuals, or even whether they might be the offspring of animals from two or more different parts of the species' range. Why is this important? If we sequence the genome of a cobra from Indonesia, but cobras in India have evolved different venom genes because of different evolutionary pressures, then we won't know that until we get some cobras from India. Taxonomic conclusions drawn from Boa constrictor gene sequences on GenBank are dubious because of the ambiguous origins of many of these specimens. The primary reasons to sequence a whole genome are subtly different from the reasons to sequence individual genes, and scientists doing these tasks have different questions. But, we should be cautious about inferring too much from the genome sequence of a single individual of any species.↩
4 Right now if you're a human you can actually get your whole genome sequenced for less than $5000, even though the first human genome cost over $3 billion, because we've optimized the process.↩
5 It's unclear how many venom proteins were originally made in the venom gland before they became toxic, and how many were recruited to this tissue following duplication. The original cobra genome paper by Vonk et al. implies that the latter is most common, whereas subsequent work by Hargreaves et al. uses gene expression data from Leopard Gecko salivary glands to suggest the former. Reyes-Velasco et al. used the python genome and transcriptome to suggest that venom genes are recruited preferentially from genes that are expressed at low levels in most tissues but at more variable levels than average across tissues.↩
6 Of the approximately 24 gene families that code for snake venom proteins, those that produce toxins that are known to be important in prey capture (e.g., the three-finger neurotoxins) have undergone repeated duplication and selection, whereas venom components that perform ancillary functions, such as helping the snake to relocate its bitten prey, do not show high rates of duplication or selection. These rates are probably further influenced by the need to target diverse receptors in different types of prey (in snakes with broad diets), and by predator-prey co-evolutionary arms races (in snakes with narrow diets).↩
7 A recent effort by a different research group generated a tree for Caenophidia using 333 loci totaling 225,140 base pairs for each of 31 snake species, almost 80,000 of which were informative. This is a drastic improvement on the 10 loci and maximum of 5,814 base pairs of the most comprehensive previous studies, but it is still a long way from the entire genome. Incredibly, they were still unable to resolve certain difficult parts of the snake family tree.↩
ACKNOWLEDGMENTS
Thanks to JD Willson, Baloch Imrankhan, and Alison Davis Rabosky for the use of their photographs, and to Alison Davis Rabosky and Todd Castoe for providing me with information regarding genomics.
REFERENCES
Alföldi et al. 2011. The genome of the green anole lizard and a comparative analysis with birds and mammals. Nature 477:587-591 <link>
Armengaud, J., J. Trapp, O. Pible, O. Geffard, A. Chaumot, and E. M. Hartmann. 2014. Non-model organisms, a species endangered by proteogenomics. Journal of Proteomics 105:5-18 <link>
Armengaud, J., J. Trapp, O. Pible, O. Geffard, A. Chaumot, and E. M. Hartmann. 2014. Non-model organisms, a species endangered by proteogenomics. Journal of Proteomics 105:5-18 <link>
Castoe et al. 2013. The Burmese python genome reveals the molecular basis for extreme adaptation in snakes. Proceedings of the National Academy of Sciences 110:20645–20650 <link>
Cox, C. L. and A. R. D. Rabosky. 2013. Spatial and Temporal Drivers of Phenotypic Diversity in Polymorphic Snakes. The American Naturalist DOI: 10.1086/670988 <link>
Gauthier, J. A., M. Kearney, J. A. Maisano, O. Rieppel, and A. D. B. Behlke. 2012. Assembling the squamate Tree of Life: perspectives from the phenotype and the fossil record. Bulletin of the Peabody Museum of Natural History 53:3-308 <link>
Cox, C. L. and A. R. D. Rabosky. 2013. Spatial and Temporal Drivers of Phenotypic Diversity in Polymorphic Snakes. The American Naturalist DOI: 10.1086/670988 <link>
Gauthier, J. A., M. Kearney, J. A. Maisano, O. Rieppel, and A. D. B. Behlke. 2012. Assembling the squamate Tree of Life: perspectives from the phenotype and the fossil record. Bulletin of the Peabody Museum of Natural History 53:3-308 <link>
Hargreaves, A. D., M. T. Swain, M. J. Hegarty, D. W. Logan, and J. F. Mulley. 2014. Restriction and recruitment-gene duplication and the origin and evolution of snake venom toxins. Genome Biology & Evolution 6:2088-2095 <link>
Hargreaves, A. D., M. T. Swain, D. W. Logan, and J. F. Mulley. 2014. Testing the Toxicofera: Comparative transcriptomics casts doubt on the single, early evolution of the reptile venom system. Toxicon. DOI:10.1016/j.toxicon.2014.10.004 <link>
Jarvis et al. 2014. Whole-genome analyses resolve early branches in the tree of life of modern birds. Science 346:1320-1331 <link>
Losos, J., D. M. Hillis, and H. W. Greene. 2012. Who speaks with a forked tongue? Science 338:1428-1429 <link>
Mackessy, S. P. and L. M. Baxter. 2006. Bioweapons synthesis and storage: The venom gland of front-fanged snakes. Zoologischer Anzeiger 245:147-159 <link>
Hargreaves, A. D., M. T. Swain, D. W. Logan, and J. F. Mulley. 2014. Testing the Toxicofera: Comparative transcriptomics casts doubt on the single, early evolution of the reptile venom system. Toxicon. DOI:10.1016/j.toxicon.2014.10.004 <link>
Jarvis et al. 2014. Whole-genome analyses resolve early branches in the tree of life of modern birds. Science 346:1320-1331 <link>
Losos, J., D. M. Hillis, and H. W. Greene. 2012. Who speaks with a forked tongue? Science 338:1428-1429 <link>
Mackessy, S. P. and L. M. Baxter. 2006. Bioweapons synthesis and storage: The venom gland of front-fanged snakes. Zoologischer Anzeiger 245:147-159 <link>
Pyron, R. A., C. R. Hendry, V. M. Chou, E. M. Lemmon, A. R. Lemmon, and F. T. Burbrink. 2014. Effectiveness of phylogenomic data and coalescent species-tree methods for resolving difficult nodes in the phylogeny of advanced snakes (Serpentes: Caenophidia). Mol. Phylogenet. Evol. 81:221-231 <link>
Reyes-Velasco, J., D. C. Card, A. Andrew, K. J. Shaney, R. H. Adams, D. R. Schield, N. R. Casewell, S. P. Mackessy, and T. A. Castoe. 2014. Expression of venom gene homologs in diverse python tissues suggests a new model for the evolution of snake venom. Molecular Biology and Evolution <link>
Schweitzer, M. H. 2011. Soft tissue preservation in terrestrial Mesozoic vertebrates. Annual Review of Earth and Planetary Sciences 39:187-216 <link>
Schweitzer, M. H. 2011. Soft tissue preservation in terrestrial Mesozoic vertebrates. Annual Review of Earth and Planetary Sciences 39:187-216 <link>
Vonk et al. 2013. The king cobra genome reveals dynamic gene evolution and adaptation in the snake venom system. Proceedings of the National Academy of Sciences 110:20651–20656 <link>
Yadav, S. P. 2007. The wholeness in suffix -omics, -omes, and the Word Om. Journal of Biomolecular Techniques 18:277 <link>
Zelanis, A. and A. Keiji Tashima. 2014. Unraveling snake venom complexity with ‘omics’ approaches: challenges and perspectives. Toxicon <link>
Zelanis, A. and A. Keiji Tashima. 2014. Unraveling snake venom complexity with ‘omics’ approaches: challenges and perspectives. Toxicon <link>
Life is Short, but Snakes are Long by Andrew M. Durso is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License.

